Cynthia Wolberger's group at the Johns Hopkins University solved a structure critical to understanding the role of ubiquitination in DNA repair and other pathways. A key event in the DNA double-strand break response is the synthesis of polyubiquitin chains consisting of the C-terminus of one ubiquitin joined to lysine 63 (K63) of the next. These chains recruit protein complexes involved in DNA repair, and must be cleared from damage sites once repair is complete. The deubiquitinating enzyme, OTUB1, inhibits synthesis of K63 polyubiquitin chains by UBC13, a ubiquitin conjugating enzyme, via a novel non-catalytic mechanism in which OTUB1 binds directly to UBC13~Ub, a covalent reaction intermediate. Inhibition requires the OTUB1 N-terminus. In addition, OTUB1 first must bind a free ubiquitin monomer, which allosterically converts OTUB1 into an inhibitor. The structure of OTUB1 bound to both an allosteric ubiquitin effector (ubiquitin aldehyde) and a UBC13~Ub covalent complex showed that the globular catalytic domain of OTUB1 binds to UBC13 while the N-terminus of OTUB1 forms an alpha-helix that contacts the ubiquitin. The structure and accompanying functional studies showed that the resulting OTUB1/ubiquitin/UBC13~Ub complex interferes with multiple aspects of the ubiquitination reaction, thereby preventing synthesis of K63 polyubiquitin. Since OTUB1 inhibits other ubiquitin conjugating enzymes in addition to UBC13, these studies open the door to further investigations of how OTUB1 regulates other ubiquitination pathways. Given the large number of both deubiquitinating and ubiquitin conjugating enzymes in the human genome, it is likely that the principles that emerged from this study will apply to the mechanisms of regulation of other ubiquitin conjugation pathways.
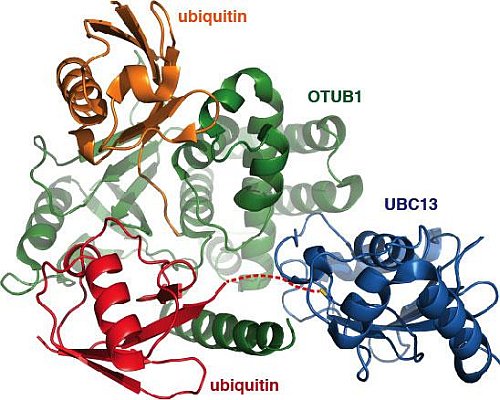 |
Figure: Structure formed between OTUB1, ubiquitin, and a UBC13~ubiquitin covalent complex. |
Citation:
Wiener, R, Zhang, X, Wang, T, Wolberger, C. The mechanism of OTUB1-mediated
inhibition of ubiquitination, Nature 483, 618-622 (2012).